헤파란황산프로테오글리칸
[ heparin sulfate proteoglycan , ~黃酸~ ]
GlcAβ1→ 3Galβ1→3Galβ1→ 4Xyl를 가교구조로 하여 헤파란황산이 핵심단백질 내의 세린에 O-글리코시드결합을 하고 있는 복합당질의 총칭. 세포 표면, 기저막 성분으로 세포외 기질에 보편적으로 존재한다. 섬유모세포 증식인자와 결합하고 기질에 저장되며 그 활성을 제어한다. 대형 헤파란황산 프로테오글리칸, 파루칸은 기저막구조의 형성에 관여하며 신장, 시냅스기저막, 근육섬유막에서 작용한다.
[네이버 지식백과] 헤파란황산프로테오글리칸 [heparin sulfate proteoglycan, ~黃酸~] (생명과학대사전, 초판 2008., 개정판 2014., 강영희)
헤파란황산
[ heparan sulfate , ~黃酸 ]
분자량 0.5~7×10-4. D-글루쿠론산, D-글루코사민의 2당단위반복구조에 N-황산화, O-황산화수식을 받은 글리코사미노글리칸. 2당단위의 황산함량은 0.4~2.0이며, L-이즈론산을 미량 함유한다. 인터류킨 3 등과 결합하며, 증식인자 활성조절인자로서 작용한다. 분해효소의 결손에 의한 유전적 대사이상증으로 헐러(Hurler)증후군 등이 알려져 있다.
[네이버 지식백과] 헤파란황산 [heparan sulfate, ~黃酸] (생명과학대사전, 초판 2008., 개정판 2014., 강영희)

SO3- termination
Heparan sulfate
From Wikipedia, the free encyclopedia
Heparan sulfate (HS) is a linear polysaccharide found in all animal tissues.[1] It occurs as a proteoglycan (HSPG, i.e. Heparan Sulfate ProteoGlycan) in which two or three HS chains are attached in close proximity to cell surface or extracellular matrix proteins.[2][3] It is in this form that HS binds to a variety of protein ligands, including Wnt,[4][5] and regulates a wide range of biological activities, including developmental processes, angiogenesis, blood coagulation, abolishing detachment activity by GrB (Granzyme B),[6] and tumour metastasis. HS has also been shown to serve as cellular receptor for a number of viruses, including the respiratory syncytial virus.[7] One study suggests that cellular heparan sulfate has a role in SARS-CoV-2 Infection, particularly when the virus attaches with ACE2.[8]
Contents
- 1Proteoglycans
- 2Structure and differences from heparin
- 3Biosynthesis
- 4Ligand binding
- 5Heparan sulfate analogue
- 6References
Proteoglycans[edit]
The major cell membrane HSPGs are the transmembrane syndecans and the glycosylphosphatidylinositol (GPI) anchored glypicans.[9][10] Other minor forms of membrane HSPG include betaglycan[11] and the V-3 isoform of CD44 present on keratinocytes and activated monocytes.[12]
In the extracellular matrix, especially basement membranes, the multi-domain perlecan, agrin and collagen XVIII core proteins are the main HS-bearing species.
Structure and differences from heparin[edit]
Heparan sulfate is a member of the glycosaminoglycan family of carbohydrates and is very closely related in structure to heparin. Heparin, commonly known as an anticoagulant, is a highly sulfated form of HS which, in contrast to HS, is mainly found in mast cell secretory granules.[13] Both consist of a variably sulfated repeating disaccharide unit. The main disaccharide units that occur in heparan sulfate and heparin are shown below.
The most common disaccharide unit within heparan sulfate is composed of a glucuronic acid (GlcA) linked to N-acetylglucosamine (GlcNAc), typically making up around 50% of the total disaccharide units. Compare this to heparin, where IdoA(2S)-GlcNS(6S) makes up 85% of heparins from beef lung and about 75% of those from porcine intestinal mucosa. Problems arise when defining hybrid GAGs that contain both 'heparin-like' and 'HS-like' structures. It has been suggested that a GAG should qualify as heparin only if its content of N-sulfate groups largely exceeds that of N-acetyl groups and the concentration of O-sulfate groups exceeds those of N-sulfate.[14]
Not shown below are the rare disaccharides containing a 3-O-sulfated glucosamine (GlcNS(3S,6S) or a free amine group (GlcNH3+). Under physiological conditions the ester and amide sulfate groups are deprotonated and attract positively charged counterions to form a salt. It is in this form that HS is thought to exist at the cell surface.
-

- GlcA-GlcNAc
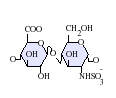
- GlcA-GlcNS

- IdoA-GlcNS
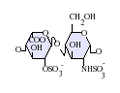
- IdoA(2S)-GlcNS
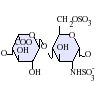
- IdoA-GlcNS(6S)
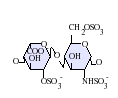
- IdoA(2S)-GlcNS(6S)
Abbreviations[edit]
- GlcA = β-D-glucuronic acid
- IdoA = α-L-iduronic acid
- IdoA(2S) = 2-O-sulfo-α-L-iduronic acid
- GlcNAc = 2-deoxy-2-acetamido-α-D-glucopyranosyl
- GlcNS = 2-deoxy-2-sulfamido-α-D-glucopyranosyl
- GlcNS(6S) = 2-deoxy-2-sulfamido-α-D-glucopyranosyl-6-O-sulfate
Biosynthesis[edit]
Many different cell types produce HS chains with many different primary structures. Therefore, there is a great deal of variability in the way HS chains are synthesised, producing structural diversity encompassed by the term "heparanome" - which defines the full range of primary structures produced by a particular cell, tissue or organism.[15] However, essential to the formation of HS regardless of primary sequence is a range of biosynthetic enzymes. These enzymes consist of multiple glycosyltransferases, sulfotransferases and an epimerase. These same enzymes also synthesize heparin.
In the 1980s, Jeffrey Esko was the first to isolate and characterize animal cell mutants altered in the assembly of heparan sulfate.[16] Many of these enzymes have now been purified, molecularly cloned and their expression patterns studied. From this and early work on the fundamental stages of HS/heparin biosynthesis using a mouse mastocytoma cell free system a lot is known about the order of enzyme reactions and specificity.[17]
Chain initiation[edit]
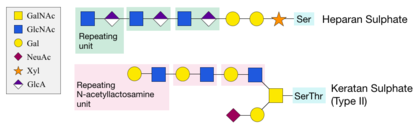
Structures of heparan sulphate and keratan sulphate, formed by the addition of xylose or GalNAc sugars, respectively, onto serine and threonine residues of proteins.
HS synthesis initiates with the transfer of xylose from UDP-xylose by xylosyltransferase (XT) to specific serine residues within the protein core. Attachment of two galactose (Gal) residues by galactosyltransferases I and II (GalTI and GalTII) and glucuronic acid (GlcA) by glucuronosyltransferase I (GlcATI) completes the formation of a tetrasaccharide primer O-linked to a serine of the core-protein:
βGlcUA-(1→3)-βGal-(1→3)-βGal-(1→4)-βXyl-O-Ser.
Xylose attachment to the core protein is thought to occur in the endoplasmic reticulum (ER) with further assembly of the linkage region and the remainder of the chain occurring in the Golgi apparatus.
The pathways for HS/heparin or chondroitin sulfate (CS) and dermatan sulfate (DS) biosynthesis diverge after the formation of this common tetrasaccharide linkage structure. The next enzyme to act, GlcNAcT-I or GalNAcT-I, directs synthesis, either to HS/heparin or CS/DS, respectively.
Chain elongation[edit]
After attachment of the first N-acetylglucosamine (GlcNAc) residue, elongation of the tetrasacchride linker is continued by the stepwise addition of GlcA and GlcNAc residues. These are transferred from their respective UDP-sugar nucleotides. This is carried out by one or more related enzymes whose genes are members of the exostoses (EXT) gene family of tumour suppressors.
Mutations at the EXT1-3 gene loci in humans lead to an inability of cells to produce HS and to the development of the disease Multiple Hereditary Exostoses (MHE). MHE is characterized by cartilage-capped tumours, known as osteochondromas or exostoses, which develop primarily on the long bones of affected individuals from early childhood until puberty.[18]
Chain modification[edit]
As an HS chain polymerises, it undergoes a series of modification reactions carried out by four classes of sulfotransferases and an epimerase. The availability of the sulfate donor PAPS is crucial to the activity of the sulfotransferases.[19][20]
N-deacetylation/N-sulfation[edit]
The first polymer modification is the N-deacetylation/N-sulfation of GlcNAc residues into GlcNS. This is a prerequisite for all subsequent modification reactions, and is carried out by one or more members of a family of four GlcNAc N-deacetylase/N-sulfotransferase enzymes (NDSTs). In early studies, it was shown that modifying enzymes could recognize and act on any N-acetylated residue in the forming polymer.[21] Therefore, the modification of GlcNAc residues should occur randomly throughout the chain. However, in HS, N-sulfated residues are mainly grouped together and separated by regions of N-acetylation where GlcNAc remains unmodified.
There are four isoforms of NDST (NDST1–4). Both N-deacetylase and N-sulfotransferase activities are present in all NDST-isoforms but they differ significantly in their enzymatic activities.[22]
Generation of GlcNH2[edit]
Due to the N-deacetylase and N-sulfotransferase being carried out by the same enzyme N-sulfation is normally tightly coupled to N-acetylation. GlcNH2 residues resulting from apparent uncoupling of the two activities have been found in heparin and some species of HS.[23]
Epimerisation and 2-O-sulfation[edit]
Epimerisation is catalysed by one enzyme, the GlcA C5 epimerase or heparosan-N-sulfate-glucuronate 5-epimerase (EC 5.1.3.17). This enzyme epimerises GlcA to iduronic acid (IdoA). Substrate recognition requires that the GlcN residue linked to the non-reducing side of a potential GlcA target be N-sulfated. Uronosyl-2-O-sulfotransferase (2OST) sulfates the resulting IdoA residues.
6-O-sulfation[edit]
Three glucosaminyl 6-O-transferases (6OSTs) have been identified that result in the formation of GlcNS(6S) adjacent to sulfated or non-sulfated IdoA. GlcNAc(6S) is also found in mature HS chains.
3-O-sulfation[edit]
Currently seven glucosaminyl 3-O-sulfotransferases (3OSTs, HS3STs) are known to exist in mammals (eight in zebrafish).[24][25] The 3OST enzymes create a number of possible 3-O-sulfated disaccharides, including GlcA-GlcNS(3S±6S) (modified by HS3ST1 and HS3ST5), IdoA(2S)-GlcNH2(3S±6S)(modified by HS3ST3A1, HS3ST3B1, HS3ST5 and HS3ST6) and GlcA/IdoA(2S)-GlcNS(3S) (modified by HS3ST2 and HS3ST4).[26][27][28][29] As with all other HS sulfotransferases, the 3OSTs use 3'-phosphoadenosine-5'-phosphosulfate (PAPS) as a sulfate donor. Despite being the largest family of HS modification enzymes, the 3OSTs produce the rarest HS modification, the 3-O-sulfation of specific glucosamine residues at the C3-OH moiety.[30]
The 3OSTs are divided into two functional subcategories, those that generate an antithrombin III binding site (HS3ST1 and HS3ST5) and those that generate a herpes simplex virus 1 glycoprotein D (HSV-1 gD) binding site (HS3ST2, HS3ST3A1, HS3ST3B1, HS3ST4, HS3ST5 and HS3ST6).[26][27][28][29][31][32][33][34][35][36][37] As the 3OSTs are the largest family of HS modification enzymes and their actions are rate-limiting, substrate specific and produce rare modifications, it has been hypothesized that 3OST modified HS plays an important regulatory role in biological processes.[29][32] It has been demonstrated that 3-O-sulfation can enhance the binding of Wnt to the glypican and may play a role in regulating Wnt in cancer.[5][10]
Ligand binding[edit]
Heparan sulfate binds with a large number of extracellular proteins. These are often collectively called the “heparin interactome” or "heparin-binding proteins", because they are isolated by affinity chromatography on the related polysaccharide heparin, though the term “heparan sulfate interactome” is more correct. The functions of heparan sulfate binding proteins ranges from extracellular matrix components, to enzymes and coagulation factors, and most growth factors, cytokines, chemokines and morphogens [38] The laboratory of Dr. Mitchell Ho at the NCI isolated the HS20 human monoclonal antibody with high affinity for heparan sulfate by phage display.[39] The antibody binds heparan sulfate, not chondroitin sulfate.[5] The binding of HS20 to heparan sulfate requires sulfation at both the C2 position and C6 position. HS20 blocks the Wnt binding on heparan sulfate[5] and also inhibits infectious entry of pathogenic JC polyomavirus.[40]
Interferon-γ[edit]
The cell surface receptor binding region of Interferon-γ overlaps with the HS binding region, near the protein's C-terminal. Binding of HS blocks the receptor binding site and as a result, protein-HS complexes are inactive.[41]
Wnt[edit]
Glypican-3 (GPC3) interacts with both Wnt and Frizzled to form a complex and triggers downstream signaling.[4][10] It has been experimentally established that Wnt recognizes a heparan sulfate modif on GPC3, which contains IdoA2S and GlcNS6S, and that the 3-O-sulfation in GlcNS6S3S enhances the binding of Wnt to the glypican.[5]
The HS-binding properties of a number of other proteins are also being studied:
- Antithrombin III
- Fibroblast Growth Factors
- Hepatocyte Growth Factor
- Interleukin-8
- Vascular Endothelial Growth Factor
- Wnt/Wingless
- Endostatin
Heparan sulfate analogue[edit]
Main article: Heparan sulfate analogue
Heparan sulfate analogues are thought to display identical properties as heparan sulfate with exception of being stable in a proteolytic environment like a wound.[42][43] Because heparan sulfate is broken down in chronic wounds by heparanase, the analogues only bind sites where natural heparan sulfate is absent and cannot be broken down by any known heparanases and glycanases.[citation needed] Also the function of the heparan sulfate analogues is the same as heparan sulfate, protecting a variety of protein ligands such as growth factors and cytokines. By holding them in place, the tissue can then use the different protein ligands for proliferation.
Entactin, 엔탁틴
Nidogen-1
From Wikipedia, the free encyclopedia
Nidogen-1 (NID-1), formerly known as entactin, is a protein that in humans is encoded by the NID1 gene.[5][6] Both nidogen-1 and nidogen-2 are essential components of the basement membrane alongside other components such as type IV collagen, proteoglycans (heparan sulfate and glycosaminoglycans), laminin[7] and fibronectin.[8]
Nidogen-2
From Wikipedia, the free encyclopedia
Nidogen-2, also known as osteonidogen, is a basal lamina protein of the nidogen family. It was the second nidogen to be described after nidogen-1 (entactin). Both play key roles during late embryonic development.[5] In humans it is encoded by the NID2 gene.[6][7]
Contents
Function[edit]
Nidogen-1 is a member of the nidogen family of basement membrane glycoproteins. The protein interacts with several other components of basement membranes. Structurally it (along with perlecan) connects the networks formed by collagens and laminins to each other.[9] It may also play a role in cell interactions with the extracellular matrix.[10][11]
Clinical significance[edit]
Mutations in NID1 cause autosomal dominant Dandy–Walker malformation with occipital encephalocele (ADDWOC).[12][13]
Interactions[edit]
Nidogen-1 has been shown to interact with FBLN1.[14][15][16]
'상식+ > 공부자료' 카테고리의 다른 글
| 온도-용해도 (0) | 2021.08.23 |
|---|---|
| Heart - basement membrane (0) | 2021.08.23 |
| BASEMENT MEMBRNAE, 기저막 (0) | 2021.08.23 |
| 흉터 없는 상처 회복 가능할까 (0) | 2021.08.23 |
| [과기원은 지금] KAIST, 나노입자 표면 리간드 분자 3차원 분포 최초 규명 外 (0) | 2021.08.19 |


